Mapping the Trees’ Tree of Life: The Genetic Lineages of North American Hazelnuts
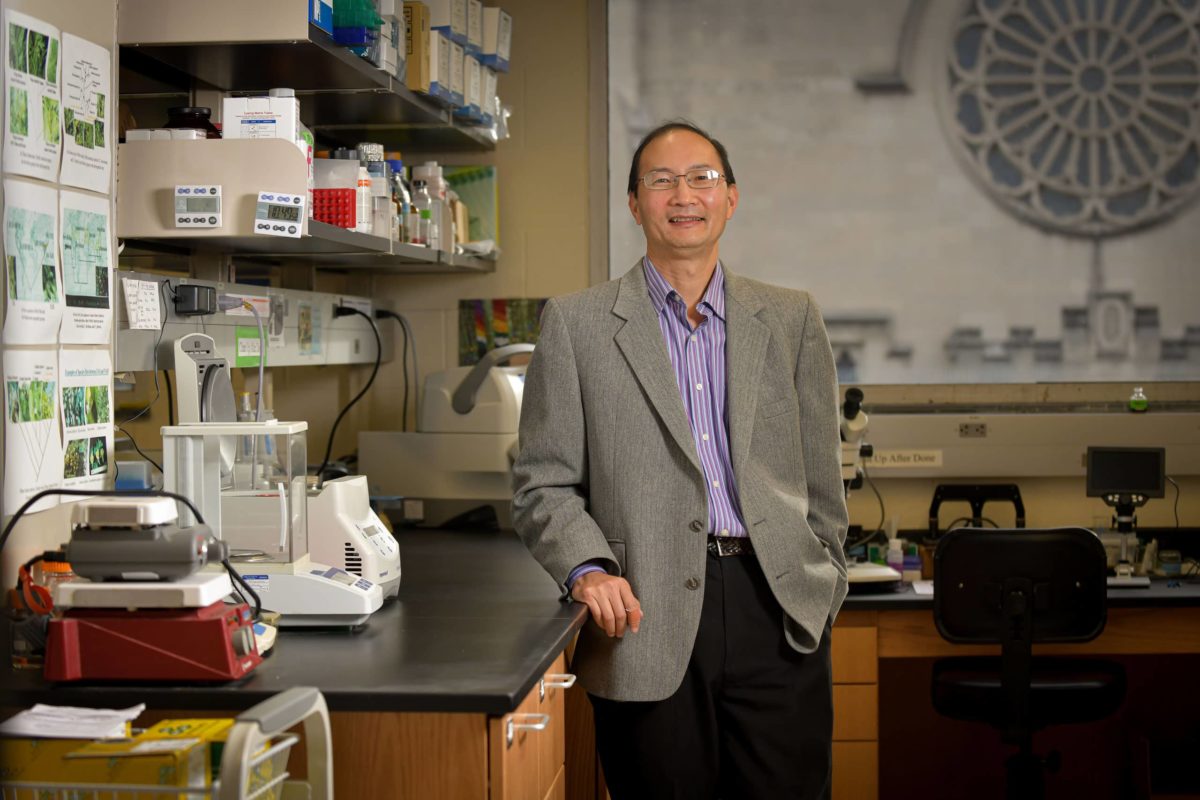
Jianhua Li, Ph.D. | Professor of Biology
Life — in all its varied forms and interactions — is one of the Earth’s most studied mysteries, one that scientists have spent centuries unraveling. Among them is botanist Dr. Jianhua Li, who is ranging through continents and (thanks to the insights of DNA) through time to uncover the branching trees of life of various, well, trees.
One of Li’s research loves is the hazelnut tree. Its several distinct species occupy the branch of the genus Corylus (which branches off the birch family, which branches off the Fagales order). The study of life is inherently intricate — indeed, labyrinthine. Li’s work aims to untangle the relationships among tree species, one study at a time.
He has been collaborating with Dr. Jianguang Xie of South China Agricultural University to examine hazelnut trees to find physical and genetic differences that set one hazelnut species apart from another. One structure, the modified leaf that covers the nuts, has a different shape from species to species: some swaddle each nut in two shrouds resembling leaves, while others wrap the nuts in a two-pronged cover that extends and fuses to form a beak shape. (The variation in this structure, immaterial though it may seem, is how the “beaked hazelnut” got its name.) Still other species use a dry, spiny structure for the same (presumably protective) function.
The hazelnut genus is one of many Li has researched over the years — and not only to puzzle over the different ways they display their fruit.
“We’re really diving into how the variation actually comes about. How have species evolved?” he says.
To begin each dive, along with cataloguing the trees’ physical appearance, he and his fellow researchers use DNA data to map out an inheritance timeline that traces which species emerged first, and where, and how it migrated and/or hybridized (interbred) with fellow hazelnut species when they occurred in the same area. Since DNA is in every cell of every living thing, this inheritance history is visible in any part of the plant — when run through the right equipment. Li usually uses leaves of various species to run these analyses, and was assisted in 2020 by Hope biochemistry and molecular biology major Tom Diaz ’21, chemistry major Yutong Zou ’22 and biology major Brittany Henkin ’20.
On behalf of the research team, in summer 2020 Dr. Xie presented their findings virtually at the Botany 2020 conference, highlighting the genetic smatterings that enabled them to trace long-ago instances of crossbreeding between North America’s two hazelnut species — the aforementioned beaked hazelnut, and the American hazelnut.
“The main thing coming out of this presentation and this paper is that, indeed, there was pretty extensive ancient hybridization between species that occur in a similar locality,” Li says.
Going back some million years in the lineages to discover this took some genetic detective work.
Plants leave clues to their lineages through three genetic pathways. One is the relatively familiar nuclear genome, the DNA in the nucleus of each cell. Second, there’s the mitochondrial genome — the DNA in the cell’s mitochondria, the famous “powerhouse” of the cell. The third is the chloroplast genome, DNA specific to the plant organelle where photosynthesis occurs. Like humans, plants inherit their nuclear genome from both parents, but their mitochondrial genome is passed down only through the maternal line (which for trees means only from the seeds, not from the pollen that fertilizes them). The chloroplast genome is also inherited only from the maternal side in most cases in flowering plants.
While the two North American species of hazelnut look different (and have been classified based on that), their chloroplast genomes are more similar than would be expected if they were completely distinct. This indicates that there was hybridization between the species at some point or points in their shared history. Their nuclear genomes, however, support the visual classification model. It’s all a bit of a puzzle.
“Now the question is: Why?” Li says. “And what might be the mechanism for that kind of difference — the difference between chloroplast and nuclear genes?” These questions outline forthcoming research avenues for him and his students.
“Maybe in the future we can do some experiments to see whether the contribution of the maternal chloroplast genome actually helps the next generation. That’s the main thing I want to put out there — that speculation,” Li says.
Beyond conjecture and the thrill of discovery, Li is also looking for real-world applications of his findings. Many of these relate to the healthcare realm. Li’s attraction to studying hazelnuts came in part from learning that their shells contain taxol, a drug first extracted from Pacific Yew tree bark in the 1960s. In 1992, the FDA approved its use in cancer treatment.
“When I choose a research project, I always want to think about both the scientific part of it and the possibility of actually applying what we find to the practical world,” Li says. “I want my students to be able to see the connection of botany with healthcare.”
While his current research does not directly investigate the genes involved in taxol production, it’s one potential future direction that could branch off the classification work he’s done to date. “It would be interesting to look at the taxol-producing pathway genes and try to figure out what’s going on,” he says. “We need to know which ones produce taxol, and how much taxol.”
In 2021, Li is turning his attention instead toward another practical concern: climate change.
“I’m thinking about what might be the things I can do to help in that area,” he says. “And, of course, one of them is carbon sequestration.” Woody plants are particularly well suited to absorbing carbon. Their lumber is latticed with lignin, a structural molecule that increases a material’s rigidity, decreases its tendency to rot, and absorbs carbon. Li has received a $5,000 grant from the Michigan Space Grant Consortium, matched by funds from Hope, to research this.
He has combed through the online database of genes already known to be involved with lignin production — there are 15 to 20 of them — and has worked with the Arbor Biosciences company in designing a probe to “fish out” these genes in his hazelnut tree and shrub samples, and in samples of other plant groups with both tree and shrub species (such as willows, dogwood and legumes). He hopes to find out if there are differences between the tree and shrub species in terms of which genes they use to spur lignin production in their tissues.
In the 2020 fall semester, Tori King ’21 and Lukas Bruxvoort ’21, student researchers in Li’s lab, extracted genomic DNA from over 60 leaf samples, quantified the amount of DNA, and sent them to Arbor Biosciences for genomic library construction and sequencing of the genes. Once they have the DNA sequences, they will use bioinformatics computer programs to compare them across the species included in the project. It’s a convoluted process, but well worth the effort for the insights gained into one more twig on the sprawling tree of life.
It’s also a process that benefits from connecting with scientists around the globe. Hazelnuts, like many plants, can be found on multiple continents — which offers additional challenge and opportunity in the quest.
“With these kinds of questions, you have to collaborate with people all over the world,” Li says. “If we are not going to do that, then any study from any country anywhere may be limited in terms of sampling. We want to be as collaborative as possible; otherwise, every study is very much just a snapshot.”